High Performance Affinity Chromatography
Quick Start
This page accompanies the paper, in Analyst: Quantifying Non-Specific Interactions via Liquid Chromatography by Seishi Shimizu, Steven Abbott, Katarzyna Adamska and Adam Voelkel, Analyst, 2019,144, 1632-1641, which is linked to the app.
HPAC data have been fitted by many ad hoc formulae. Here we implement the analysis discussed in the paper which uses Kirkwood-Buff integrals to yield fundamental insights into the interactions.
High Performance Affinity Chromatography
The interaction of a ligand with a protein bound to a support material can be changed with different concentrations of a co-solute which can be something as simple as a salt. The classic analysis has used the idea that co-solute has a binding constant to the protein and that the binding affects the binding of the ligand. The correct approach makes no hypothesis of the specifics, allowing many different possible interactions to change the interactions of the ligand with the polymer. Instead of the classic log(k) [retention factor] versus log(concentration), we plot ln(k) versus concentration, and provide a fitting function to the data. The slope of this fitted curve at any point gives us the KBI value, ΔGu2 along with the excess number, N, of the co-solute (salt) molecules at the given concentration.
The datasets provided are all from a paper by Slama et al1 containing an extensive set of analyses of HPAC data. The analyses in the paper are the "classical" type which the KBI analysis makes redundant.
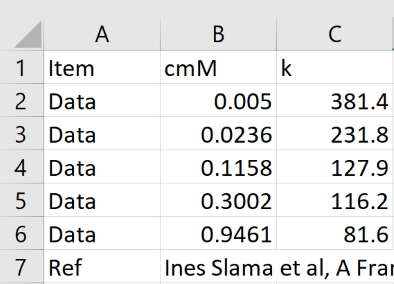 To analyse your own dataset, provide it as an Excel .csv file as per this image. The column headers and row names are case sensitive so have to be exact. The Ref row is optional - whatever is in the second column appears in the Ref line in the app. The Data rows (up to 10) hold the cosolute concentration (mM) and k. You can also type values in to the table by hand.
To analyse your own dataset, provide it as an Excel .csv file as per this image. The column headers and row names are case sensitive so have to be exact. The Ref row is optional - whatever is in the second column appears in the Ref line in the app. The Data rows (up to 10) hold the cosolute concentration (mM) and k. You can also type values in to the table by hand.
Codes for the examples: Asp-Chy-NaP = Aspartame-α-Chymotrypsin-NaPhosphate; Med-Gly-NaP = Medetomide-α-1-Glycoprotein-NaPhosphate; DDT-VAN-NaC = D-Dansyl Tryptophan-Vancomycin-NaCitrate; DDT-VAN-NaCl = D-Dansyl Tryptophan-Vancomycin-NaChloride; DDV-VAN-NaC = D-Dansyl Valine-Vancomycin-NaCitrate; DDT-VAN-NaCl = D-Dansyl Valine-Vancomycin-NaChloride; PRO-GLY-NaP = Propranolol-α- 1-Glycoprotein-NaPhosphate; BUP-GLY-NaP = Bupranolol-α- 1-Glycoprotein-NaPhosphate; IBU-GLY-NaP = Ibuprofen-α- 1-Glycoprotein-NaPhosphate; NBP-BSA-NaP = N-Benzoyl Phenylalanine-Bovine Serum Albumin-NaPhosphate; TRY-HSA-NaP = L-Tryptophan-Human Serum Albumin-NaPhosphate
1Ines Slama, Corinne Ravelet, Catherine Grosset, Anne Ravel, Annick Villet, Edwige Nicolle, and Eric Peyrin, A Framework Based on the Extended Wyman Concept for Analyzing the Salt Effects on the Solute Retention in High-Performance Affinity Chromatography, Anal. Chem. 2002, 74, 282-287

